The following page describes all the settings for the parameters calculations.
- Calculations settings
- \(\lambda_z\) calculation settings
- Parameters to compute
- Acceptance criteria settings
- Ratios for NCA parameters
Calculations
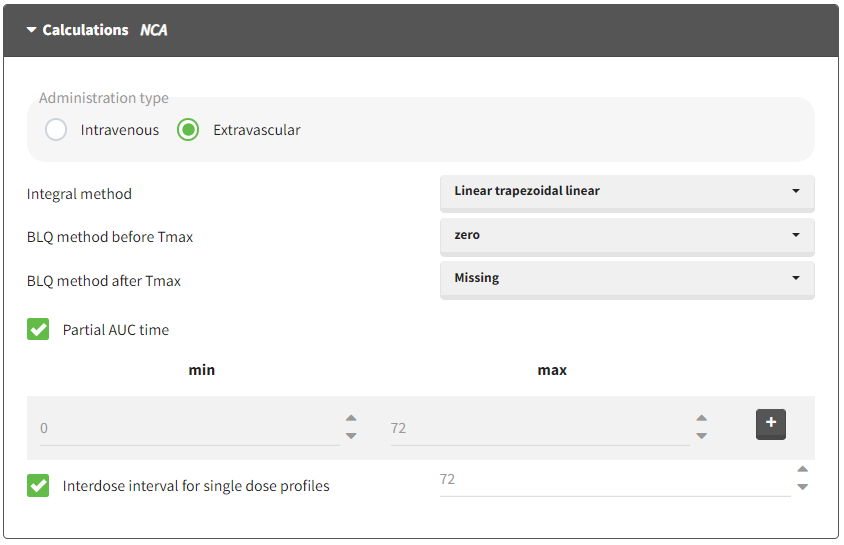
These settings corresponds to the settings impacting the calculation of the NCA parameters.
- Administration type: Intravenous or Extravascular. This defines the drug type of administration. IV bolus and IV infusions must be set as “intravenous”.
- Integral method: Method for AUC and AUMC calculation and interpolation. Trapezoidal refers to formula to calculate the AUC and Interpolation refers to the formula to add an additional point in case of partial AUC with a time point not originally present in the data set. See the Calculation rules for details on the formulas.
- Linear Trapezoidal Linear (Linear trapezoidal, linear interpolation): For AUC calculation linear formula. For interpolation linear formula.
- Linear Log Trapezoidal (Linear/log trapezoidal, linear/log interpolation): For AUC, linear before Cmax and log after Cmax. For interpolation, linear before Cmax and log after.
- Linear Up Log Down (LinUpLogDown trapezoidal, LinUpLogDown interpolation): For AUC, linear if concentration is going up or is stable, log if going down. For interpolation, linear if concentration is going up or is stable, log if going down.
- Linear Trapezoidal linear/log (Linear trapezoidal, linear/log interpolation): For AUC, linear formula. For interpolation, linear before Cmax, log after Cmax. If several Cmax, the first one is used.
For all methods, if an observation value is less than or equal to zero, the program defaults to the linear trapezoidal or interpolation rule for that point. Similarly, if adjacent observations values are equal to each other, the program defaults to the linear trapezoidal or interpolation rule.
- Partial AUC time: Define if the user would like to compute a partial AUC in a specific time interval. It is possible to define several partial AUC intervals, applicable to all individuals.
- BLQ before Tmax: In case of BLQ measurements, this corresponds to the method by which the BLQ data before Tmax should be replaced. Possible methods are “missing”, “LOQ”, “LOQ/2” or “0” (default value is 0).
- BLQ after Tmax: In case of BLQ measurements, this corresponds to the method by which the BLQ data after Tmax should be replaced. Possible methods are “missing”, “LOQ”, “LOQ/2” or “0” (default value is LOQ/2).
-
Interdose interval for single dose profiles: To compute steady-state parameters for single dose profiles, a value for \( \tau\) can be manually specified. The input value may be a double, under the condition that it is strictly positive. By default, the last time point of the initial single dose profile is set. This option is available with PKanalix version 2024R1 and above.
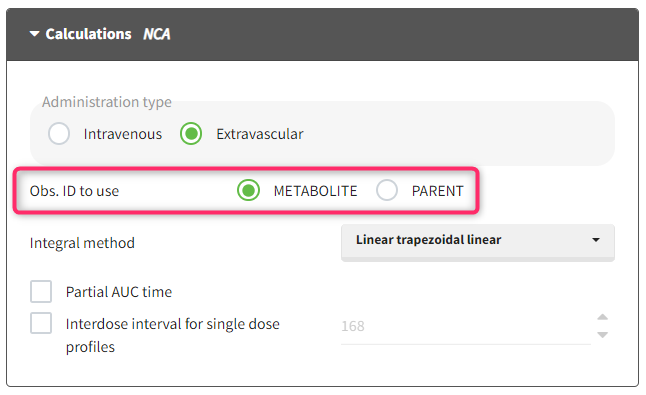
- Obs. ID to use: When several OBS ID are present in the data, an additional option to choose which Obs ID to use for perform the NCA analysis appears.
Settings impacting the \(\lambda_z\) calculation
This settings are impacting the calculation of \(\lambda_z\). See also the Check lambda_z page.
- Main rule for the \(\lambda_z\) estimation. It corresponds to the rule to define the measurements used for the \(\lambda_Z\) calculation. Possible rules are “R2”, “interval”, “points” or “adjustedR2” (called Best Fit in Winonlin) (default value is “adjustedR2”). See the calculation rules for more details.
- In case of “R2” or “adjustedR2” rule, the user has the possibility to define the maximum number of points and/or minimum time for the \(\lambda_z\) estimation. It allows to constrain the points tested for inclusion.
- In case of “Interval” rule, the user has the possibility to define the time interval to consider.
- In case of “Points” rule, the user has the possibility to define the number of points to consider.
- Weighting method used for the regression that estimates \(\lambda_z\). Possible methods are “Y”, “Y2” or “uniform” (default value is “uniform”).
Selecting NCA parameters to compute
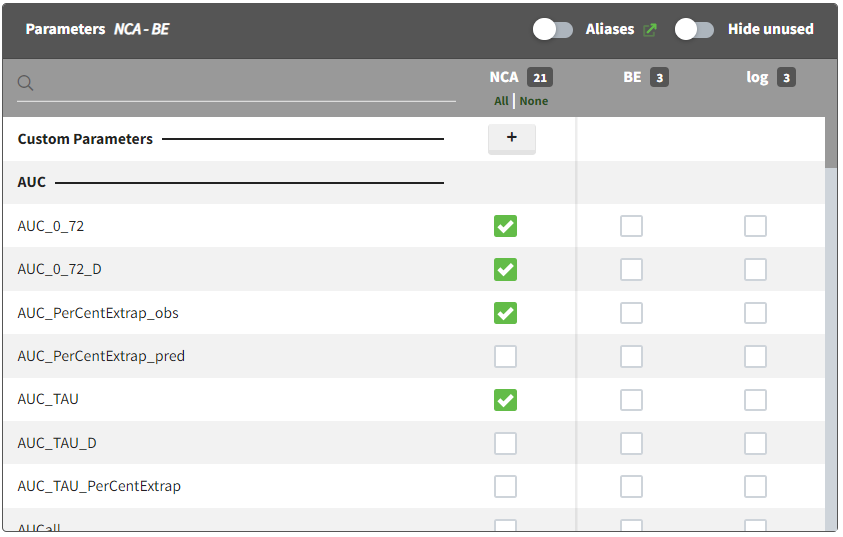
The NCA parameters to compute can be ticked in the “NCA” column. The list of NCA parameters is ordered by types of parameters. Custom NCA parameters can be defined using the “+” button.
It is recommended to always compute Rsq_adjusted, No_points_lambda_z, Lambda_z and Lambda_z_intercept as these parameters are necessary to display the terminal slope and information box in the NCA fit plot.
The list of parameters computed by default when starting a new PKanalix project can be defined in the Settings > Preferences.
Acceptance criteria
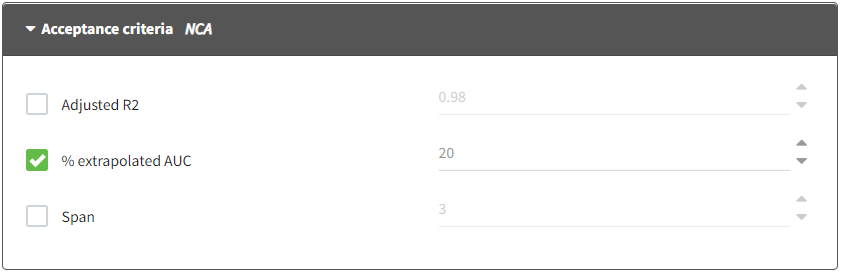
These settings corresponds to the settings impacting the acceptance criteria. When acceptance criteria are defined, flags indicating if the condition is met or not are added to the output result table. Statistics in the summary table can be calculated using only individuals who satisfy one or several acceptance criteria. The filtering option is available directly in the Summary sub-tab of the NCA results.
- Adjusted R2: It corresponds to the threshold of the adjusted R2 for the estimation of \(\lambda_z\). If activated, it will fill the value of Flag_Rsq_adjusted: if Rsq_adjusted > threshold, then Flag_Rsq_adjusted=1. The default value is .98.
- % extrapolated AUC: It corresponds to the threshold of the percentage of the total predicted AUC (or AURC for urine models) due to the extrapolation to infinity after the last measurement. If activated, it will fill the value of Flag_AUC_%Extrap_pred: if AUC_%Extrap_pred < threshold, then Flag_AUC_%Extrap_pred=1. The default value is 20%.
- Span: It corresponds to the threshold of the span. The span corresponds to the ratio of the sampling interval length for the \(\lambda_z\) calculation and the half-life. If activated, it will fill the value of Flag_Span: if Span > threshold, then Flag_Span=1. The default value is 3.
Ratios for NCA parameters
Starting from PKanalix version 2024R1 and above, ratios for each individual of NCA metrics along occasions can now be calculated. In the dedicated section “Ratios NCA” in the NCA tasks tab, ratios can be defined by clicking on the plus sign (red frame). This action prompts a window to appear, allowing you to specify the ratio in greater detail.
The Ratio definition box appears only if occasions are defined in the dataset.
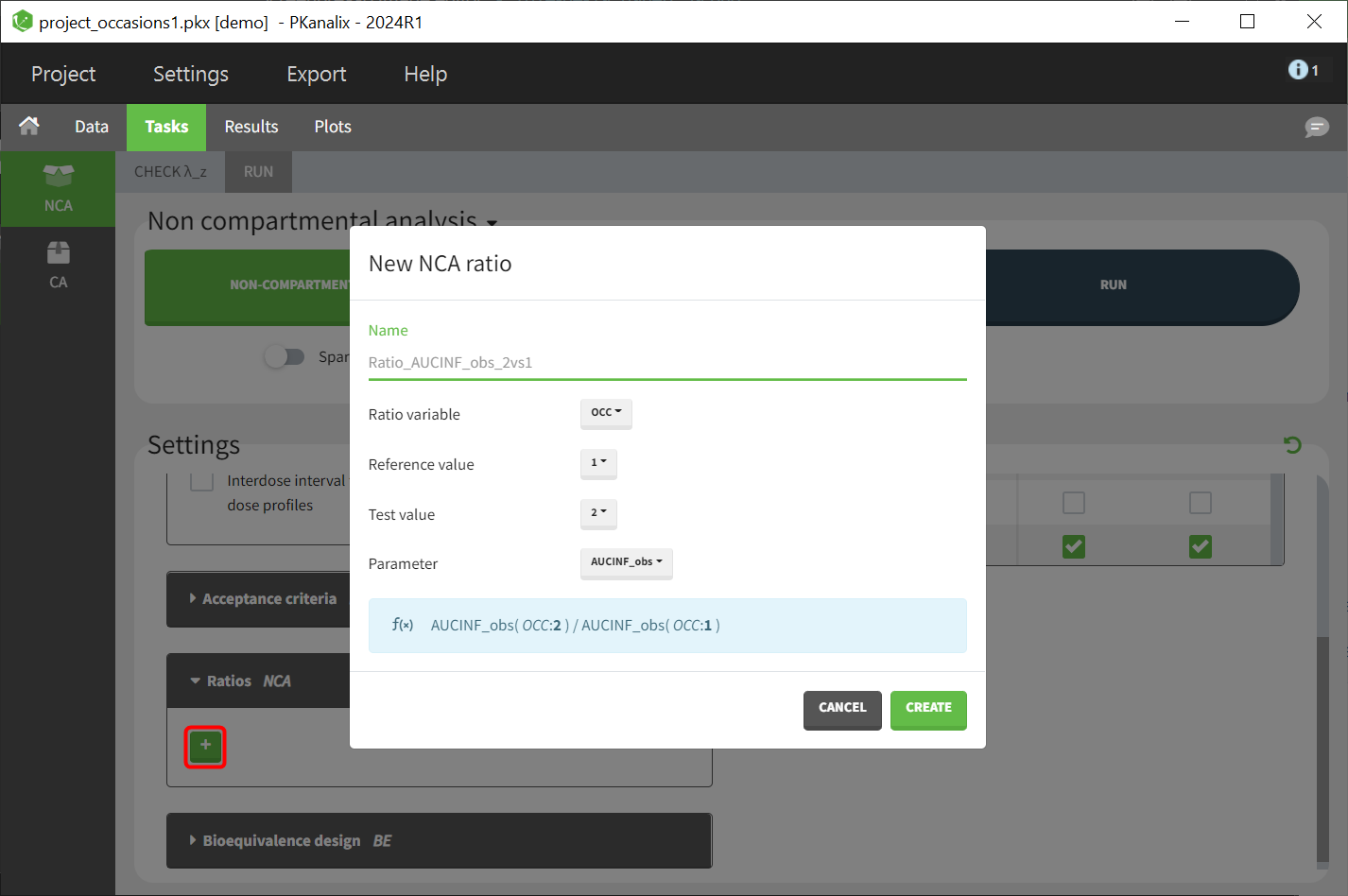
The definition is organised in several steps:
- Name: name for the new ratio parameter
- Ratio variable: Selection of the stratification variable. Variables tagged in the dataset either as occasion or as categorical variable are available for selection.
- Reference value: Selection of the reference modality (specific modality in Occasion or covariate vriable) to be used as denominator.
- Test value: Selection of test modality (specific modality in Occasion or covariate variable) to be used as numerator.
- Parameter: Selection of the NCA parameter
The formula for the specified ratio is displayed in the blue panel beneath the definition options.
Ratios can only be created using parameters that have been selected in the “NCA” column of the main parameter selection section.
For calculation details, see the Calculations rules page.
The computed parameter ratios are presented in two distinct results tables within the results tab. One table provides an individual listing, while the other offers a summary table with basic descriptive statistical metrics.