Overview of the workflow
One of the main features of PKanalix is the calculation of the parameters in the compartmental analysis framework. It find parameters of a compartmental model representing the PK dynamics for each individual using the Nelder-Mead algorithm.
NEW: PKanalix 2023 version performs individual model fit using any model with continuous outputs – loaded from the built-in library or a custom model.
Compartmental analysis workflow includes:
CA task
To access the compartmental analysis task in PKanalix use a dedicated section (left panel) in the “Tasks” tab, as in the following figure.
This task contains three sub-tabs:
- The first one called “MODEL” allows to select a model, e.g. from a PKanalix models library or your directory, edit it and match the observations from a dataset with model outputs. Complete guide to model definition is in the CA model page.
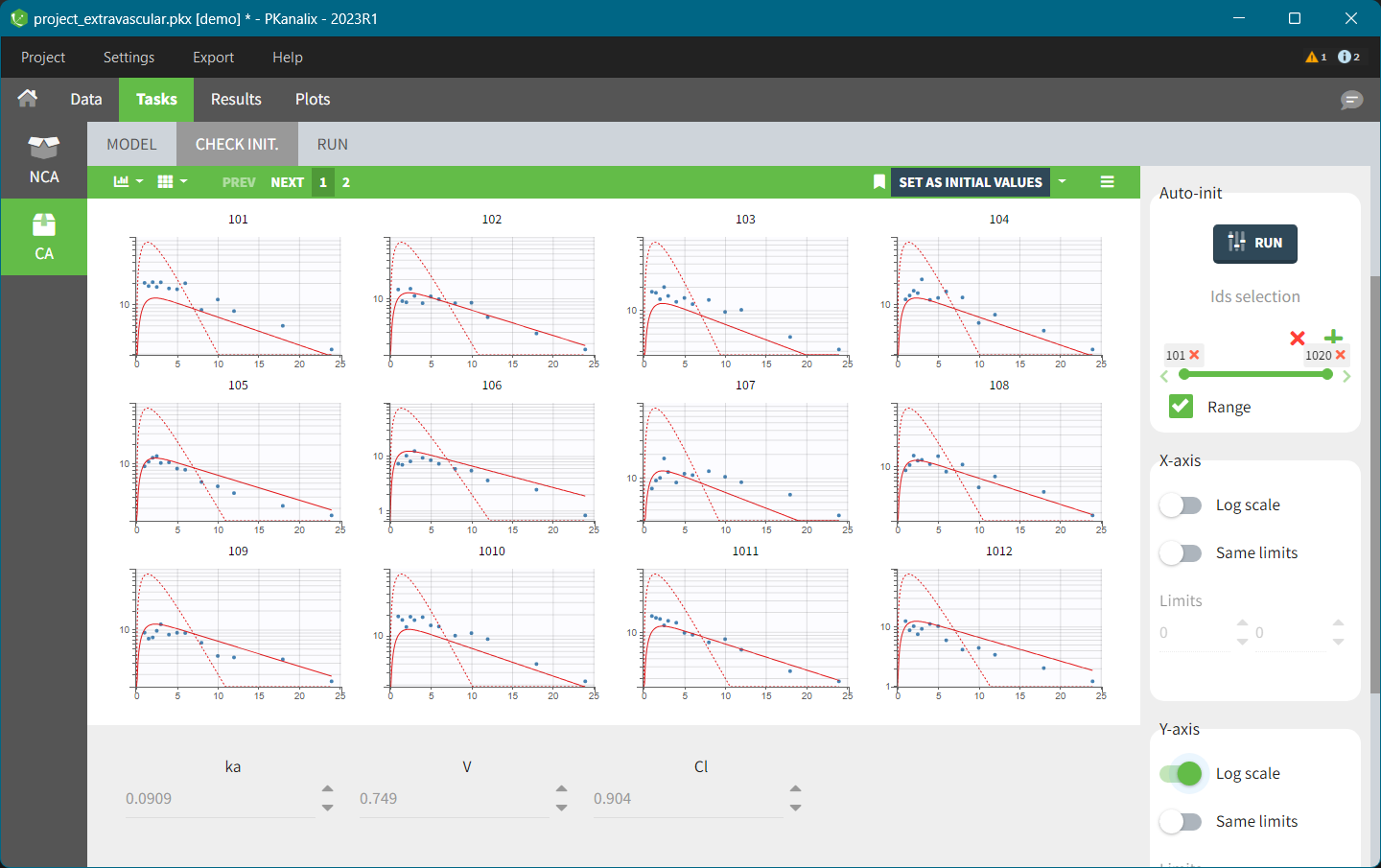
- The second one called “CHECK INIT.” shows model predictions obtained with the initial parameter values for each individual together with the data points. It helps to initialize model parameters by changing values manually or using the Auto-init function, as explained in the Initialization page.
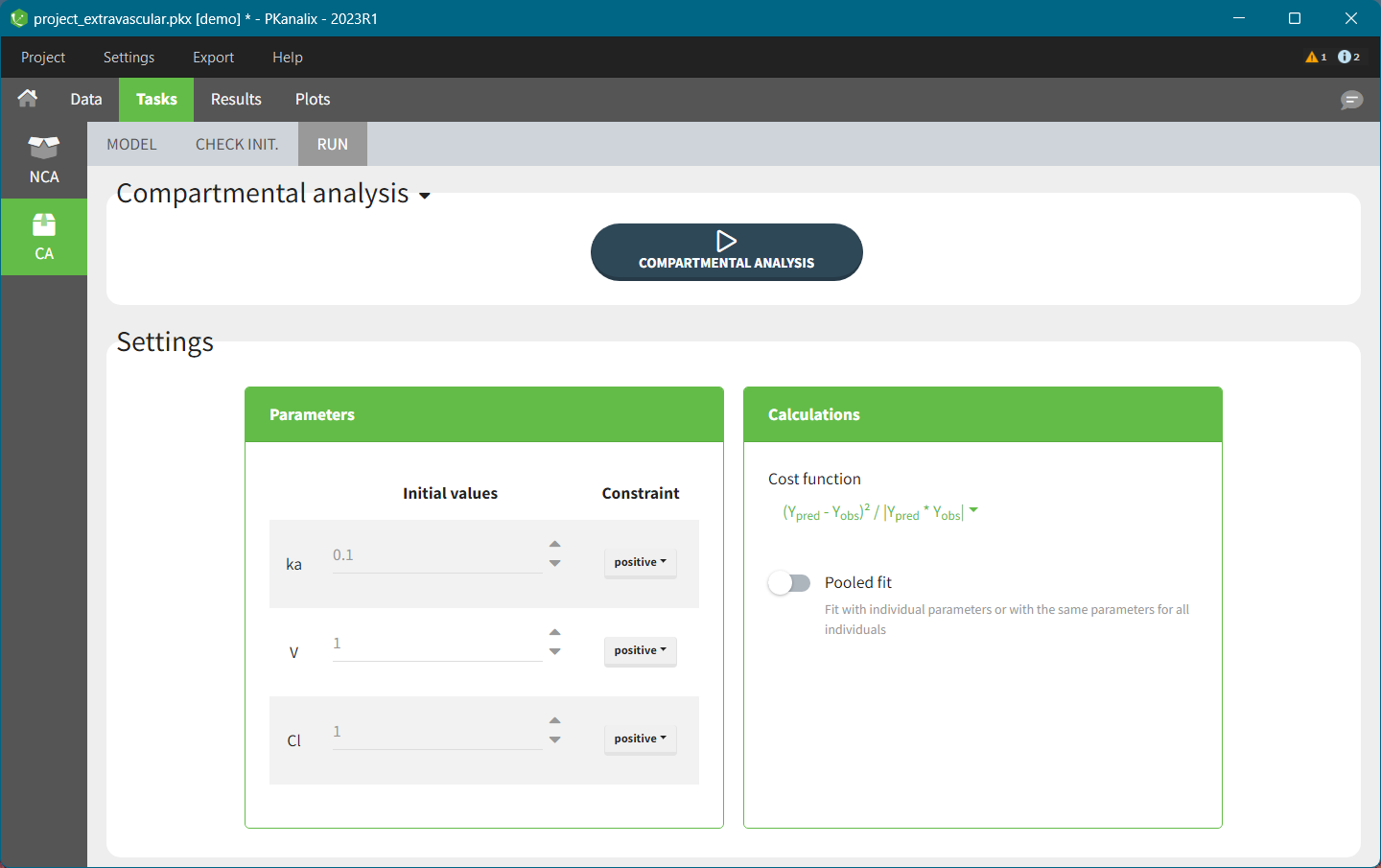
- The third one called “Run” contains a button to run the calculations and the settings for the model and the calculations. The meaning of all the settings and their default values is defined the Settings page.
CA model
In this section you specify a model. You can edit the selected model directly in the interface. The right hand side section is to match observations from a dataset with model outputs, see here for a complete guide.
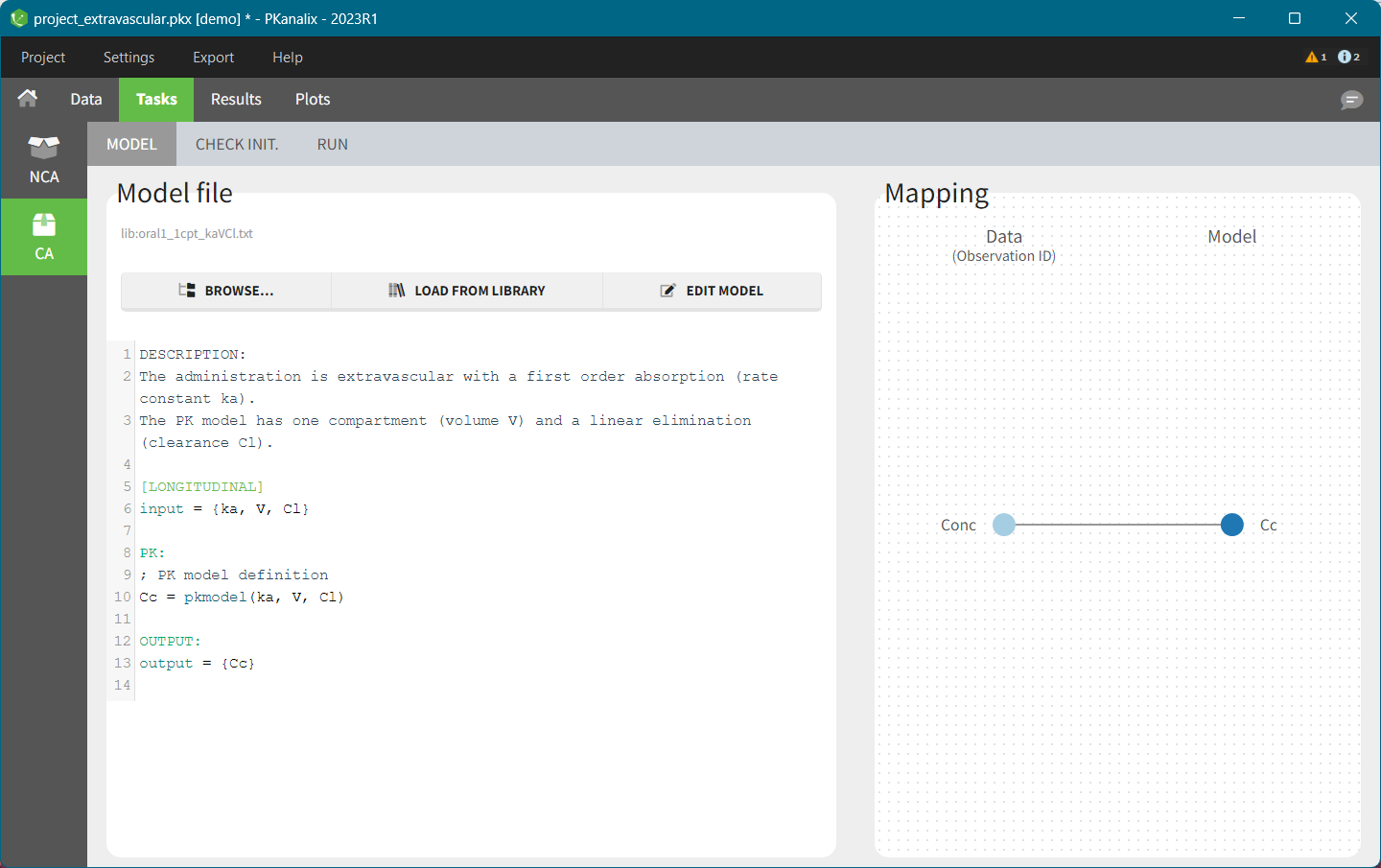
PKanalix accepts custom models – select BROWSE and load a .txt file with the model. There are also built-in libraries (PK, PK double absorption, TMDD, parent – metabolite, etc.), to facilitate the use of the software. Note that models can include multiple outputs. Models used in all MonolixSuite applications have to be in the mlxtran language. Note: For PKanalix versions before 2023, only the classical PK models, from the PK model library, are available.
CA check init.
The “Check init” tab helps to initialize model parameters. It shows the model predictions obtained with the initial model parameter values and the individual designs (doses and regressors). Each sub plot shows one individual with overlaid individual data points. This feature is very useful to find “good” initial values, as explained here.
CA results
When you run the CA task, Pkanalix shows automatically the results in the “Results” tab, see the CA Results page for a complete guide. There are three tables:
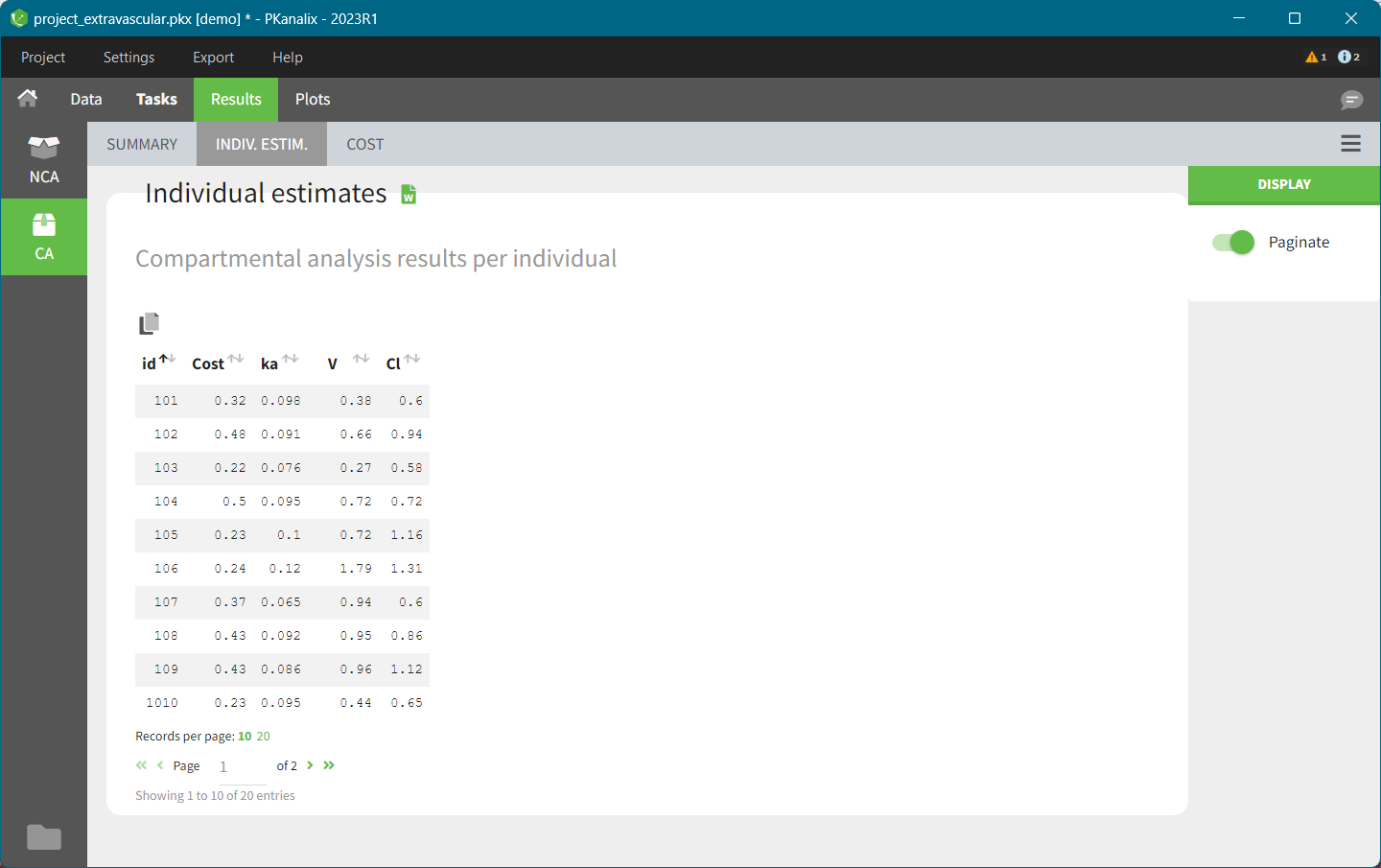
- INDIV. ESTIM contains compartmental analysis results for each individual.
- SUMMARY contains statistics on compartmental analysis results and summary calculation (described here). You can split or filter this table by covariates present in a dataset.
- COST contains cost function information.
All the computed parameters depend on the chosen model. To copy a table and paste in a word or an excel document click on the icon on the top left corner of a table.
CA plots
After running the CA task, several plots associated to the individual parameters are automatically generated and displayed In the “Plots” tab.
- Individual fits: displays model fit for each individual on a separate plot.
- Correlation between CA parameters: displays scatter plots for each pair of parameters and allows to identify correlations between parameters.
- Distribution of the CA parameters: displays the empirical distribution of the parameters and allows to analyse their distribution over the individuals.
- CA parameters w.r.t. covariates: displays the individual parameters as a function of the covariates and allows to identify correlation effects between the individual parameters and the covariates.
- Observations vs predictions displays observations from a dataset versus model predictions computed using individual parameters and is useful to detect misspecifications in the model.
CA outputs
After running the CA task, the following files are automatically generated in the project Result folder/IndividualParameters/ca :
- summary.txt contains the summary of the CA parameters calculation, in a format easily readable by a human (but not easy to parse for a computer).
- cost.txt contains the information about total cost and information criteria for a current model.
- caIndividualParametersSummary.txt contains the summary of the CA parameters in a computer friendly format useful for post-processing:
- The first column corresponds to the name of the parameters.
- The other columns correspond to the several elements describing the summary of the parameters (as explained here).
- caIndividualParameters.txt contains the CA parameters for each subject-occasion along with the covariates.
-
- The first line corresponds to the name of the parameters.
- The other lines correspond to the value of the parameters.
-
You can export the files caIndividualParametersSummary.txt and caIndividualParameters.txt in R for example using the following command
read.table("/path/to/file.txt", sep = ",", header = T)
Remark: the separator is the one defined in the user preferences. We set “,” in this example as it is the one by default.
CA export to Monolix/Simulx
Using the top menu “Export”, a PKanalix project (dataset information, model, estimated parameters) can be exported to Monolix or Simulx for further analysis, see Export to Monolix/Simulx page for a complete guide.
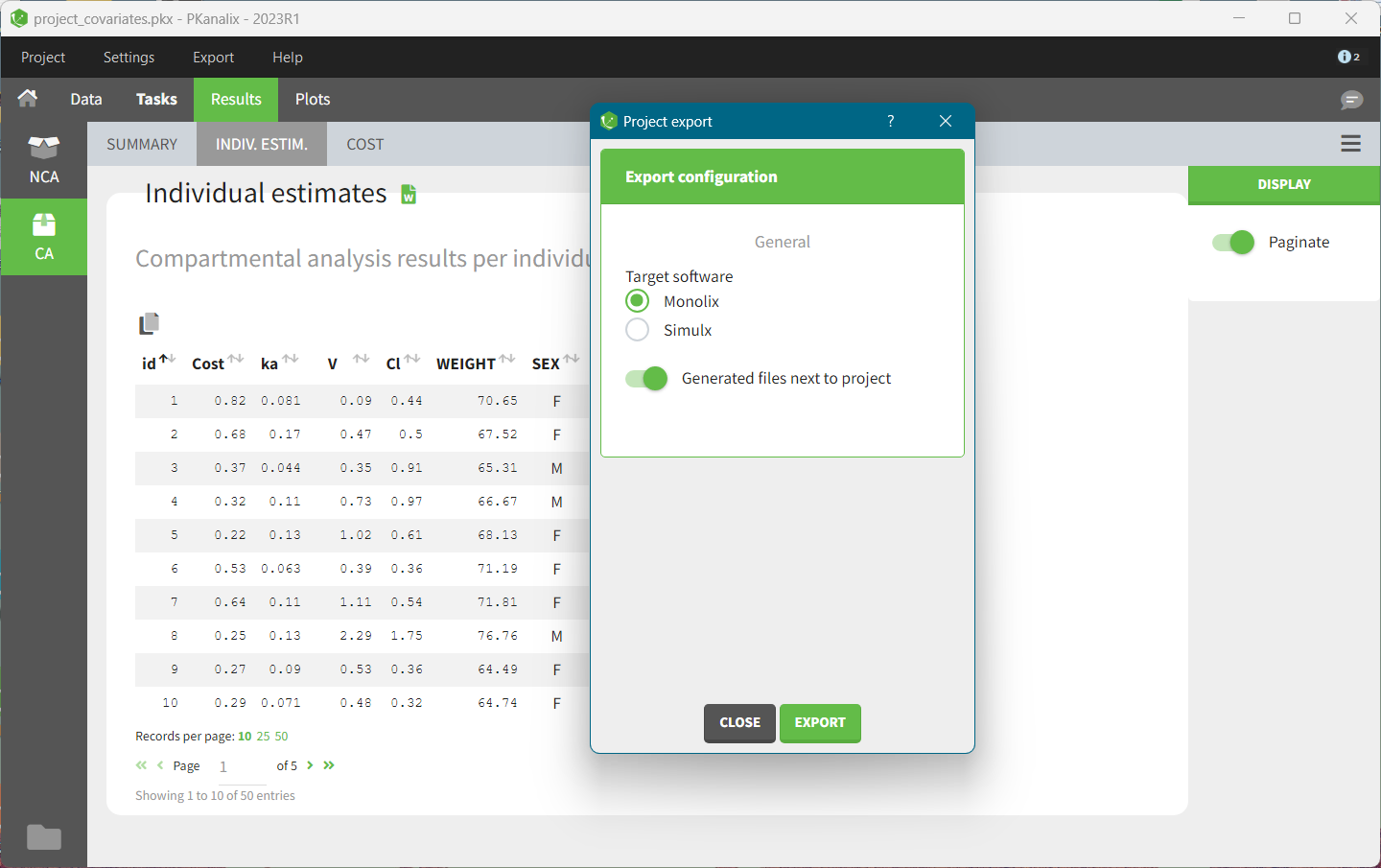
- Monolix for population modeling and further model development and diagnosis.
- Simulx to use a CA model and estimated individual parameters for simulations of new scenarios, eg. new dosing regimens.
Note: For PKanalix versions before 2023, only export to Monolix is available.
Next section: CA model