One of the main feature of PKanalix is the calculation of the parameters in the Non Compartmental Analysis framework.
NCA task
Main function of PKanalix is to calculate PK metrics (non-compartmental parameters). In the Tasks tab, you find a dedicated section “NCA” (left panel). There are two subtabs: RUN and CHECK \(\lambda_z\).
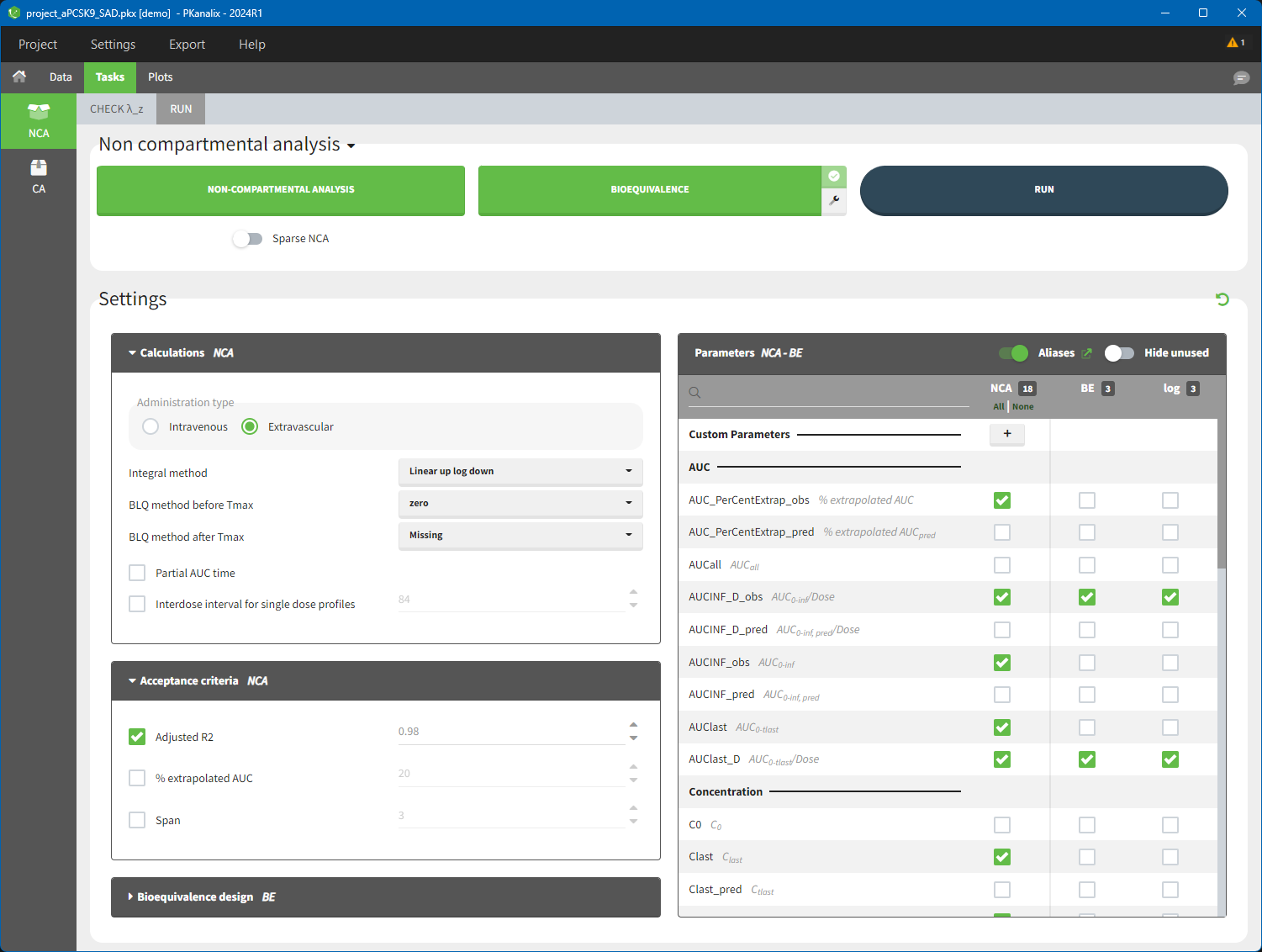
- RUN tab contains section with calculation task buttons (for non-compartmental analysis and bioequivalence) and the settings: calculation, acceptance criteria, parameters and bioequivalence design. The meaning of all the settings and their default is in a separate page here.
- CHECK \(\lambda_z\) allows to define the calculation of \(\lambda_z\) – it is a graphical preview of each individual data and the regression line, more details are here.
To calculate NCA parameters click on the button NON-COMPARTMENTAL ANALYSIS. If you click on RUN button, then PKanalix runs all selected tasks (“in green”). Results and plots are automatically generated in separate tabs.
NCA results
When you run the NCA task, results are automatically generated and displayed in the RESULTS tab. There are three tables: individual estimates, summary and points for λ_z.
Non compartmental analysis results per individual
Individual estimates of the NCA parameters are displayed in the table in the tab “INDIV. ESTIM.”
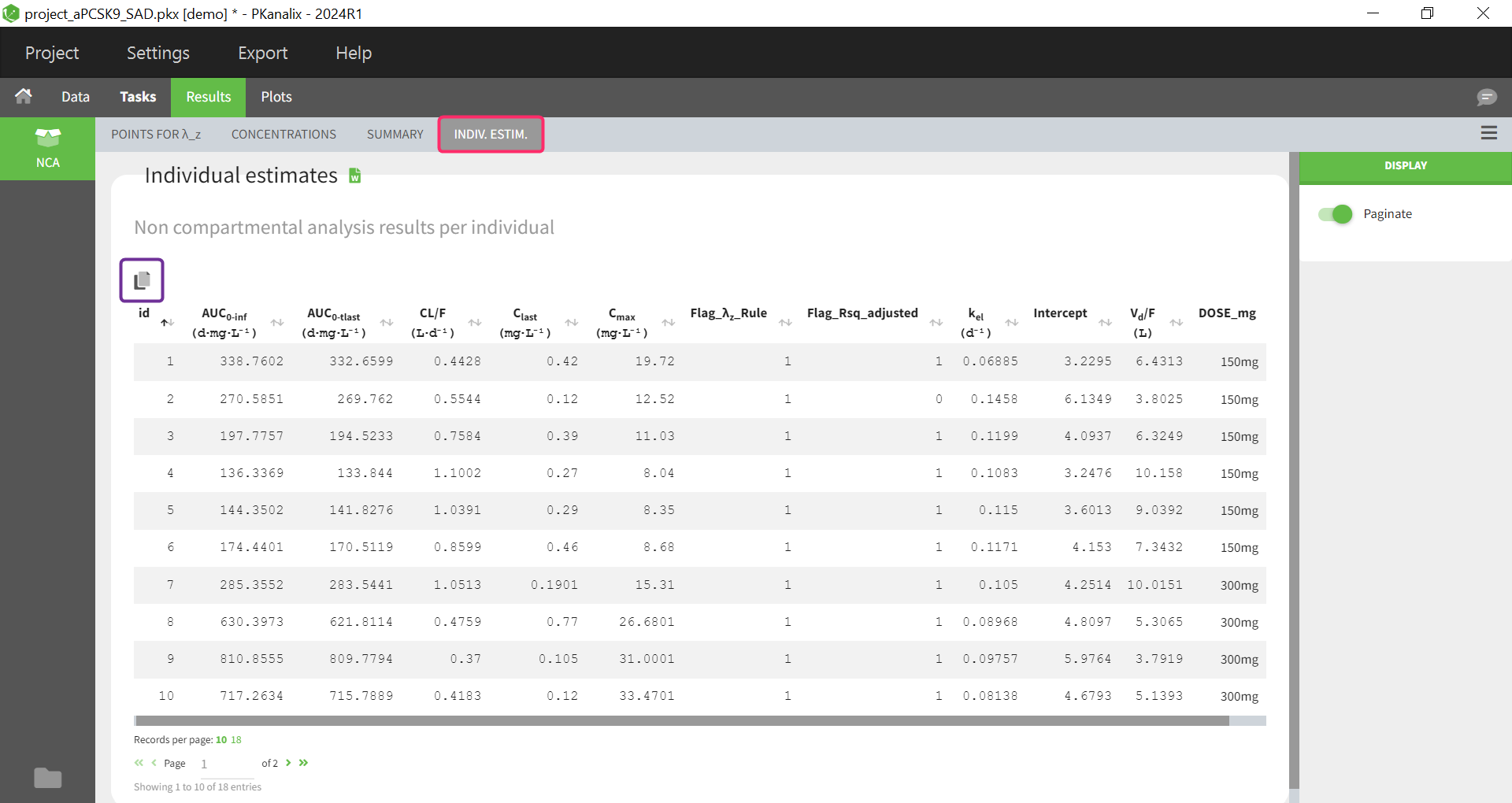
All the computed parameters depend on the type of observation and administration. Description of all the parameters is here. You can copy this table to a word or an excel file using an icon on the top right (purple frame).
In addition to generated flag variables defined by defined acceptance criteria in the NCA settings here, the Flag_\( \lambda_{z}\)_Rule column is now available since version 2024. If the inclusion/exclusion of data points in the calculation of lambda z do not follows the main rule (data points have been manually selected), the Flag_\( \lambda_{z}\)_Rule column is filled with 0, otherwise with 1.
Summary statistics on NCA results
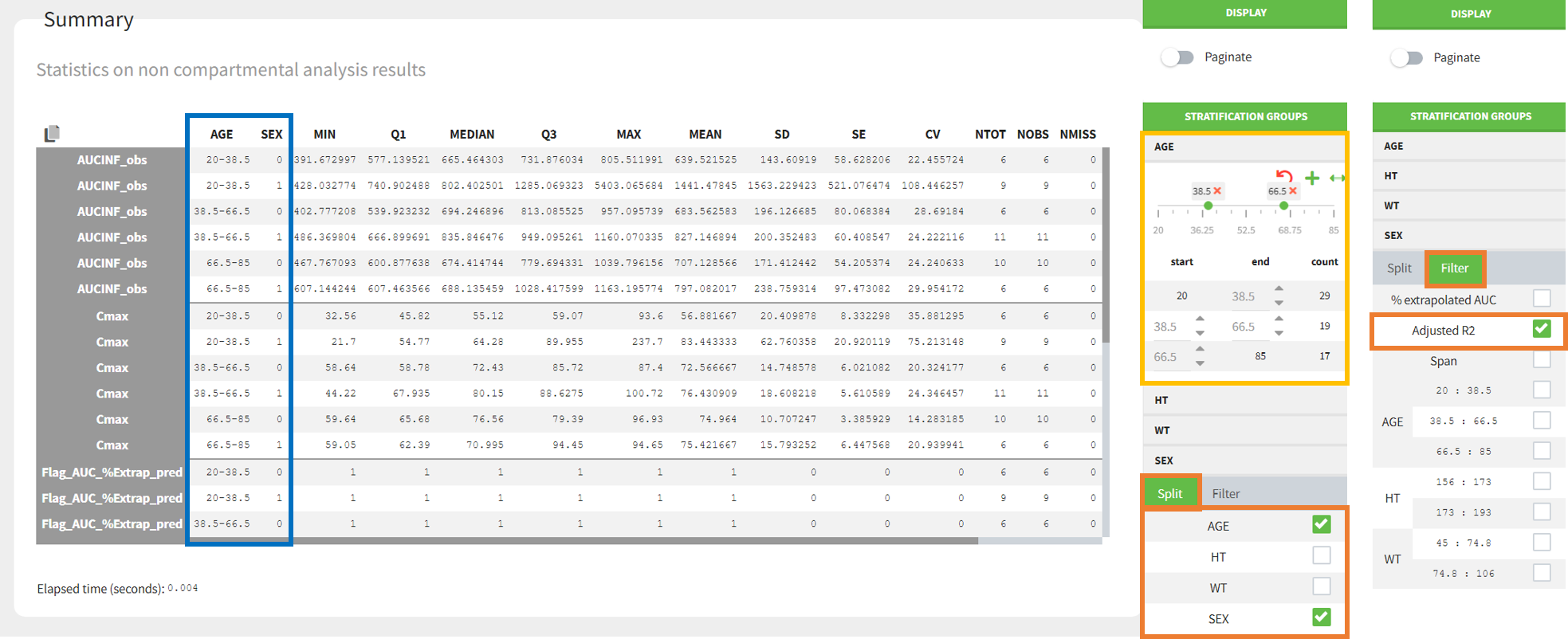
Statistics on non-compartmental parameters are displayed in the summary table in the sub-tab “SUMMARY”. Description of all the summary calculation is here.
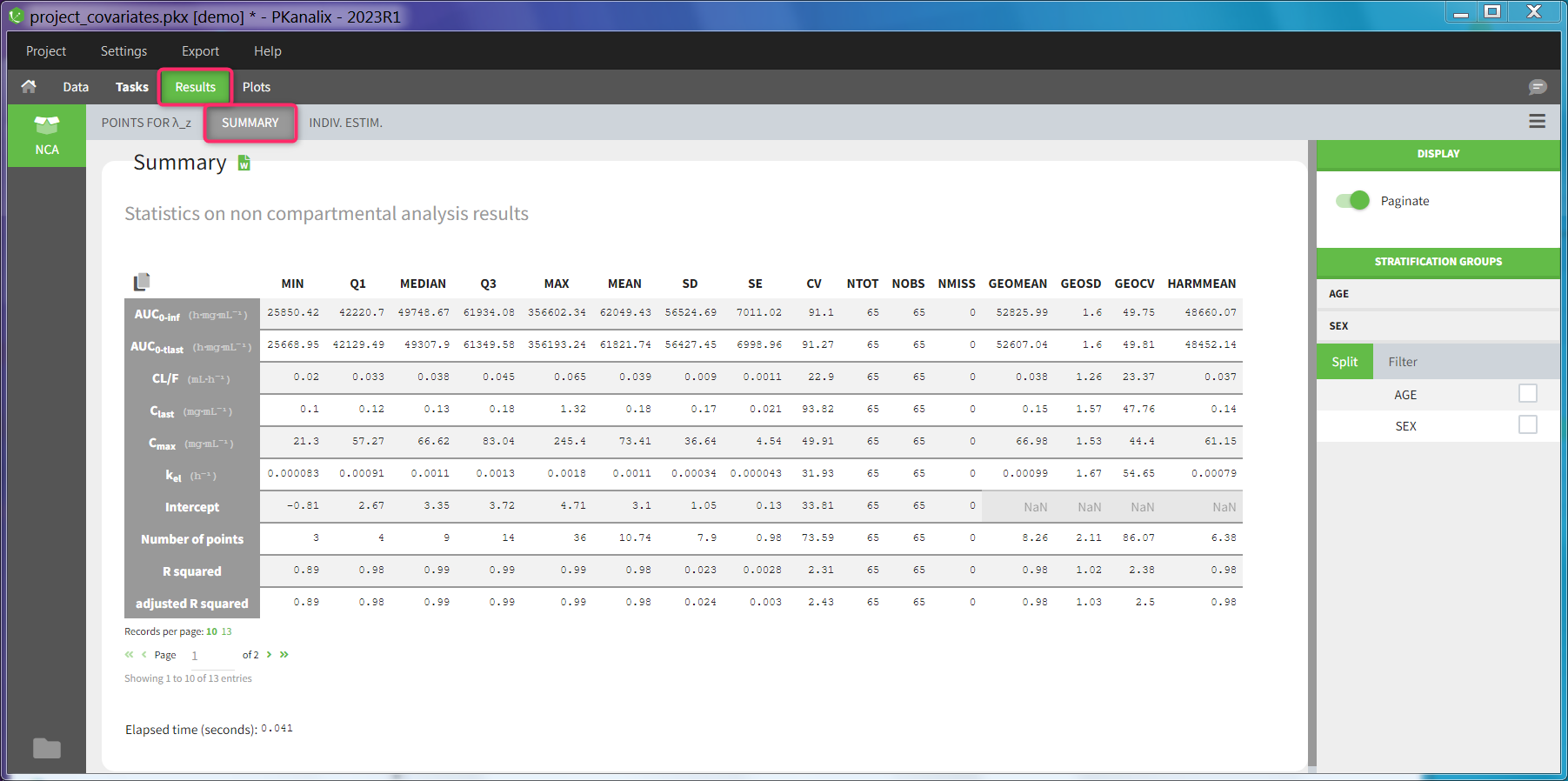
The summary table can be
- split and filtered according to the categorical and continuous covariates tagged in the data set
- filtered according to the acceptance criteria defined in the NCA settings
- Split by covariates: When you apply “split” of a table, the values of the splitting covariates are displayed in the first columns of the summary table (blue highlight). The order of these columns corresponds to the order of the clicks to setup the splitting covariates (orange highlight). You can discretized continuous covariates into groups by defining the group limits (yellow highlight), and create new categories for categorical covariates (drug and drop to merge existing categories).
It is currently not possible to split the table in several subtables (instead of splitting the rows), nor to choose the orientation of the table (NCA parameters as columns for instance). See the reporting documentation for details how these options can be applied in a report. - Filter by acceptance criteria: you can remove from the calculations in the summary table parameter values from subjects that do not meet the acceptance criteria
Read details on filtering rules
- When filtering by the “% extrapolated AUC” criterion, values from subjects that do not meet the criterion are excluded for the following parameters: [AUCINF_obs, AUCINF_pred, AUCINF_D_obs, AUCINF_D_pred, AUC_PerCentExtrap_obs, AUC_PerCentExtrap_pred, Vz_obs, Vz_pred, Vz_F_obs, Vz_F_pred, Cl_obs, Cl_pred, Cl_F_obs, Cl_F_pred, AUMCINF_obs, AUMCINF_pred, AUMC_PerCentExtrap_obs, AUMC_PerCentExtrap_pred, MRTINF_obs, MRTINF_pred, Vss_obs, Vss_pred, AURC_INF_obs, AURC_INF_pred, AURC_PerCentExtrap_obs, AURC_PerCentExtrap_pred].
- When filtering by the “Adjusted R2” or “Span” criteria, all parameters in the previous list are similarly impacted, as well as the additional following parameters: [Rsq, Rsq_adjusted, Corr_XY, Lambda_z, Lambda_z_intercept, Lambda_z_lower, Lambda_z_upper, HL_Lambda_z, Span, Clast_pred, No_points_lambda_z, AUC_PerCentBack_Ext_obs, AUC_PerCentBack_Ext_pred, Rate_last_pred].
- The following parameters are not impacted by the filtering: [AUC_T1_T2 AUC_T1_T2_D, AUC_TAU, AUC_TAU_D, AUC_TAU_PerCentExtrap, AUCall, AUClast, AUClast_D, AUMC_TAU, Accumulation_Index, C0, CLss, Cavg, Cavg_0_20, Clast, Cmax, Cmax_D, Cmin, Ctau, Ctrough, Dose, FluctuationPerCent, FluctuationPerCent_Tau, N_Samples, Swing, Swing_Tau, T0, Tau, Tlast, Tmax, Tmin, Vz]. In particular, parameters that relate to steady-state or partial AUC are not filtered, even though they rely on the lambda_z value for extrapolation when no valid observation is available at the upper end of the interval.
When you save a PKanalix project, the stratification settings of the results are saved in the result folder and are reloaded when reloading a PKanalix project. The table in the <result folder>/PKanalix/IndividualParameters/nca/ncaIndividualParametersSummary.txt takes into account the split definition. This table is generated when clicking “run” in the task tab (usually without splits, as the result tab is not yet available to define them) and also upon saving the project (with splits if defined).
Table of concentrations
With MonolixSuite version 2024, a table of concentrations is now generated after each NCA. It can either be displayed as a summary table or as a table for individual concentrations.
Summary table of concentrations
This table is displayed by default. The columns contain the concentrations of different time points, which are summarised along the rows by different descriptive metrics.
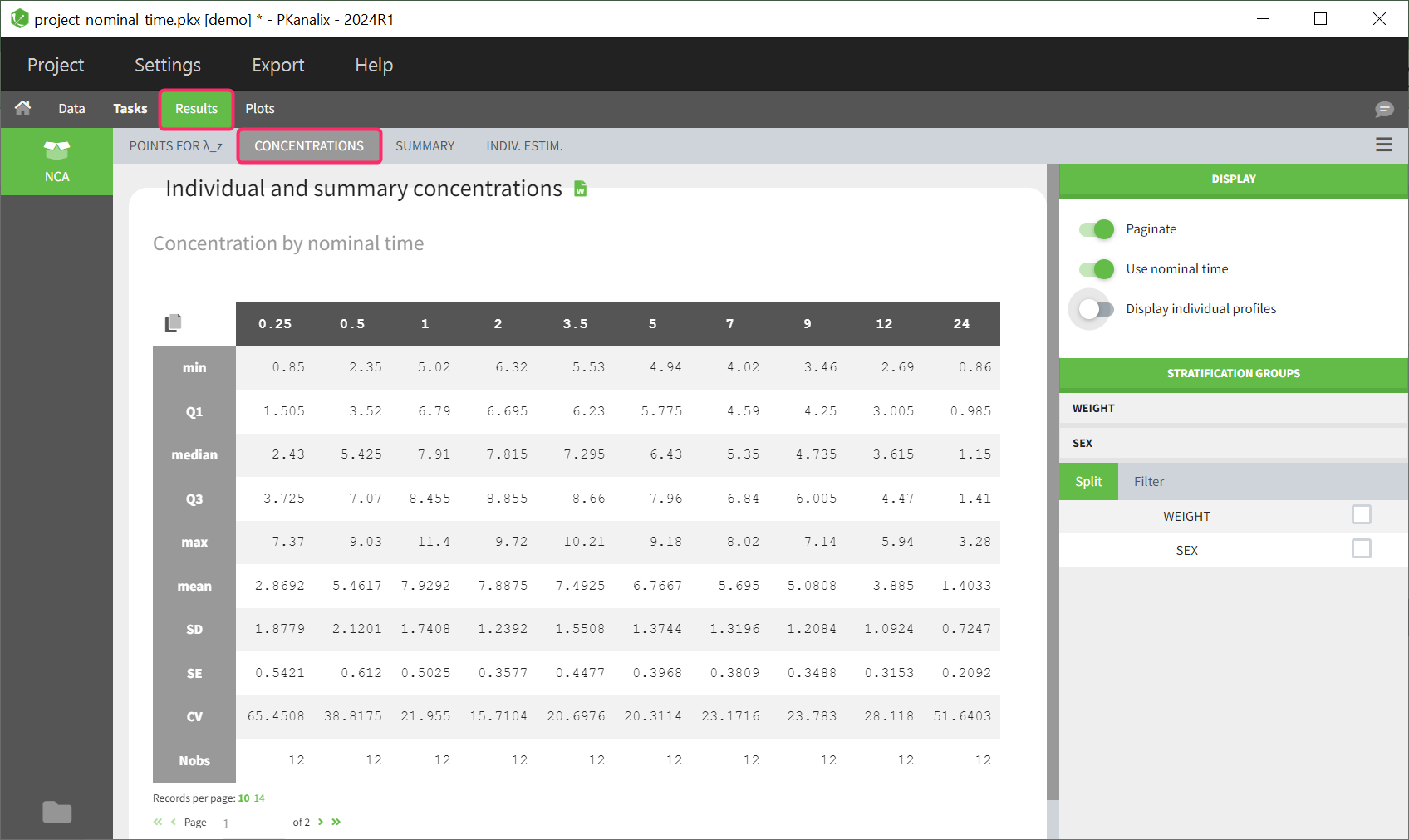
If nominal time is available in the dataset, the concentrations are automatically summarized using nominal time points. By disabling the toggle in the right panel, the table can also be displayed based on the actual time contained in the dataset.
The same stratification and filter options are available as for the NCA summary table.
The rules established in the NCA task tab for handling censored data are also considered in the calculation of the descriptive statistics.
Table of individual concentrations
This table can be displayed in the same tab by enabling the toggle “Display individual profiles” in the right-hand side panel.
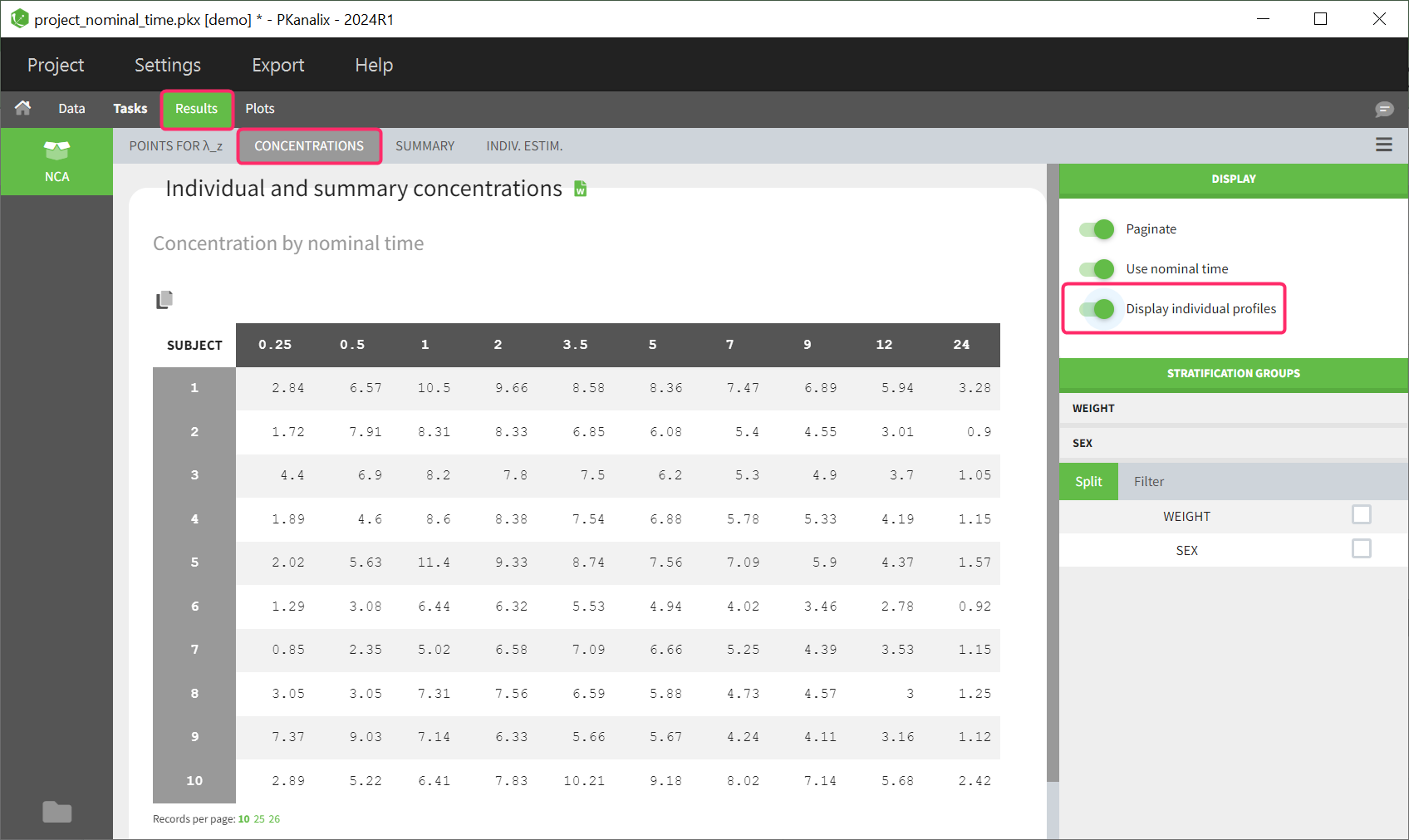 Each row in the table corresponds to an individual. The columns correspond to the different time points. There are options for stratification and filtering. Additionally, displaying the columns based on the nominal time or actual time points is also optional. Extrapolated points at dosing time are not displayed in the table as well as unused data for NCA, such as datapoints before last dose or negative values.
Each row in the table corresponds to an individual. The columns correspond to the different time points. There are options for stratification and filtering. Additionally, displaying the columns based on the nominal time or actual time points is also optional. Extrapolated points at dosing time are not displayed in the table as well as unused data for NCA, such as datapoints before last dose or negative values.
If the dataset contains left censored data, these will be displayed in the individual table of concentrations using the “<LLOQ” tag.
Ratios for NCA parameters
With MonolixSuite version 2024, parameter ratios can be calculated if required. Once this task has been performed, the results are displayed in 2 different tables in the results tab, as a summary table and as a table of individual ratios.
Summary table of ratios
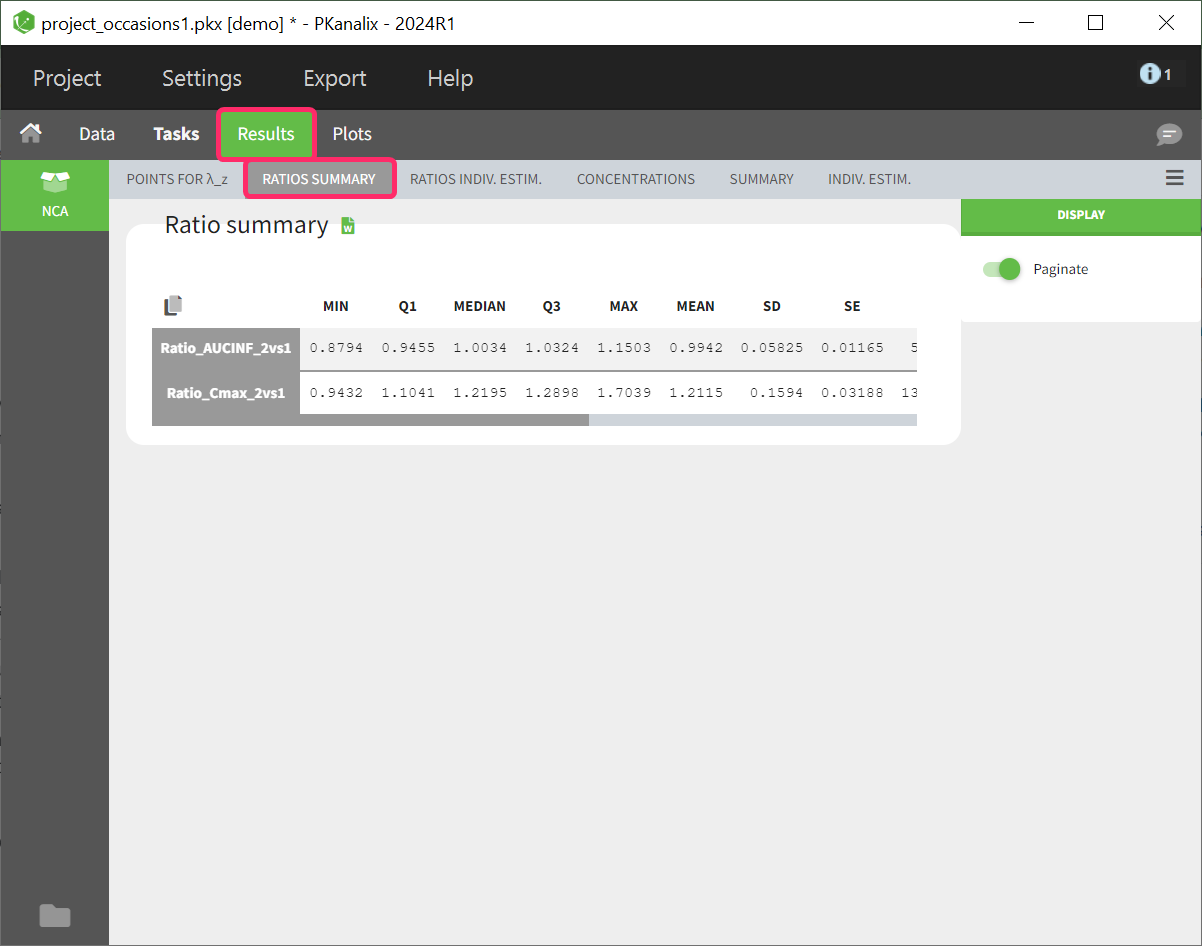
The calculated parameter ratios are presented using basic descriptive statistics. However, even if the dataset contains additional covariates or occasions, this table cannot be further stratified.
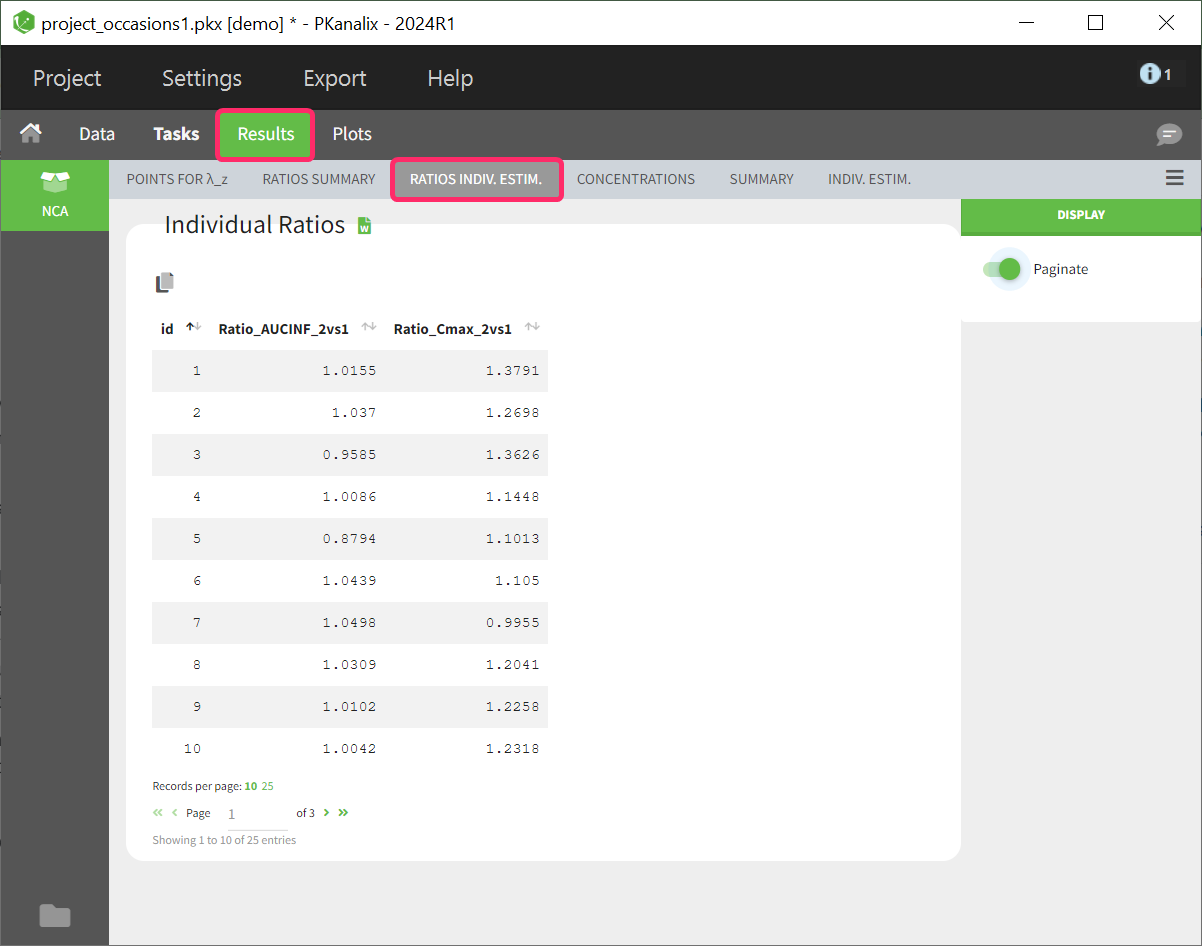
Table of ratios per individual
The parameter ratio is calculated per individual. This table shows a list sorted by subject identifier. The number of columns in the table depends on how many ratios have been defined. If a ratio couldn’t be calculated for a subject, such as due to missing values, NaN is displayed as the entry for the corresponding ratio parameter.
Points for lambda_z
In the sub-tab “POINTS FOR λ_z ” you find table of concentration points used for the terminal slope (λz) calculation.
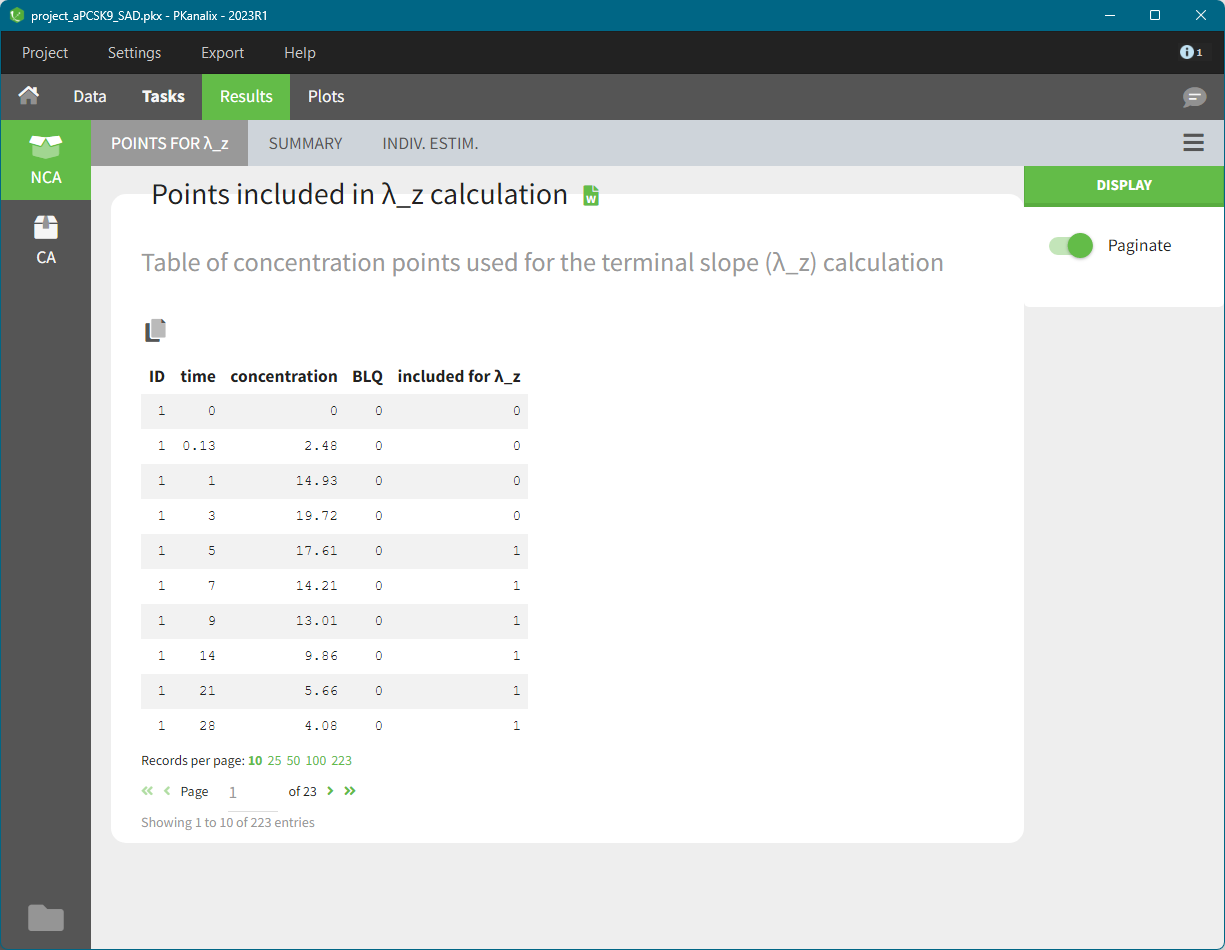
This table is displayed only if number of rows <5000. Above this, the tab appears as usual but with incomplete data. You can find the full table in the result folder: <result folder>/PKanalix/IndividualParameters/nca/pointsIncludedForLambdaZ.txt.
NCA plots
The PLOTS tab shows plot associated to the individual parameters:
- Correlation between NCA parameters: displays scatter plots for each pair of parameters. It allows to identify correlations between parameters, which can be used to see the results of your analysis and see the coherence of the parameters for each individuals.
- Distribution of the NCA parameters: shows the empirical distribution of the parameters. it allows to verify distribution of parameters over the individuals.
- NCA parameters w.r.t. covariates: displays the individual parameters as a function of the covariates. It allows to identify correlation effects between the individual parameters and the covariates.
- NCA individual fits: shows the lambdaZ regression line for each individual.
- NCA data: displays the data according to the NCA settings as spaghetti plot
NCA outputs
After running the NCA task, the following files are available in the result folder: <resultFolder>/PKanalix/IndividualParameters/nca
- Summary.txt contains the summary of the NCA parameters calculation, in a format easily readable by a human (but not easy to parse for a computer)
- ncaIndividualParametersSummary.txt contains the summary of the NCA parameters in a computer-friendly format.
- The first column corresponds to the name of the parameters
- The second column corresponds to the CDISC name of the parameters
- The other columns correspond to the several elements describing the summary of the parameters (as explained here)
- ncaIndividualParameters.txt contains the NCA parameters for each subject-occasion along with the covariates.
- The first line corresponds to the name of the parameters
- The second line corresponds to the CDISC name of the parameters
- The other lines correspond to the value of the parameters
- pointsIncludedForLambdaZ.txt contains for each individual the concentration points used for the lambda_z calculation.
- id: individual identifiers
- occ: occasions (if present). The column header corresponds the data set header of the column(s) tagged as occasion(s).
- time: time of the measurements
- concentration: concentration measurements as displayed in the NCA individual fits plot (i.e after taking into the BLQ rules)
- BLQ: if the data point is a BLQ (1) or not (0)
- includedForLambdaZ: if this data point has been used to calculate the lambdaZ (1) or not (0)
The files ncaIndividualParametersSummary.txt and ncaIndividualParameters.txt can be exported in R for example using the following command
read.table("/path/to/file.txt", sep = ",", header = T)
Remarks
- To load the individual parameters using PKanalix name as headers, your just need to skip the second line
ncaParameters = read.table("/path/to/file.txt", sep = ",", header = T); ncaParameters[-1,] # to remove the CDISC name line - To load the individual parameters using CDISC as headers, your just need to skip the second line
ncaParameters = read.table("/path/to/file.txt", sep = ",", header = T, skip = 1) - The separator is the one defined in the user preferences. We set “,” in this example as it is the one by default.